PiGx is a collection of genomics pipelines. All pipelines are easily configured with a simple sample sheet and a descriptive settings file. The result is a set of comprehensive, interactive HTML reports with interesting findings about your samples.
Wurmus R, Uyar B, Osberg B, Franke V, Gosdschan A, Wreczycka K, Ronen J, Akalin A. PiGx: Reproducible genomics analysis pipelines with GNU Guix. Gigascience. 2018 Oct 2. doi: 10.1093/gigascience/giy123. PubMed PMID: 30277498.
PiGx includes the following pipelines:
PiGx BSseq for raw fastq read data of bisulfite experiments
PiGx RNAseq for RNAseq samples
PiGx scRNAseq for single cell dropseq analysis
PiGx ChIPseq for reads from ChIPseq experiments
PiGx CRISPR (work in progress) for the analysis of sequence mutations in CRISPR-CAS9 targeted amplicon sequencing data
Here are some teaser snapshots taken from the HTML reports:
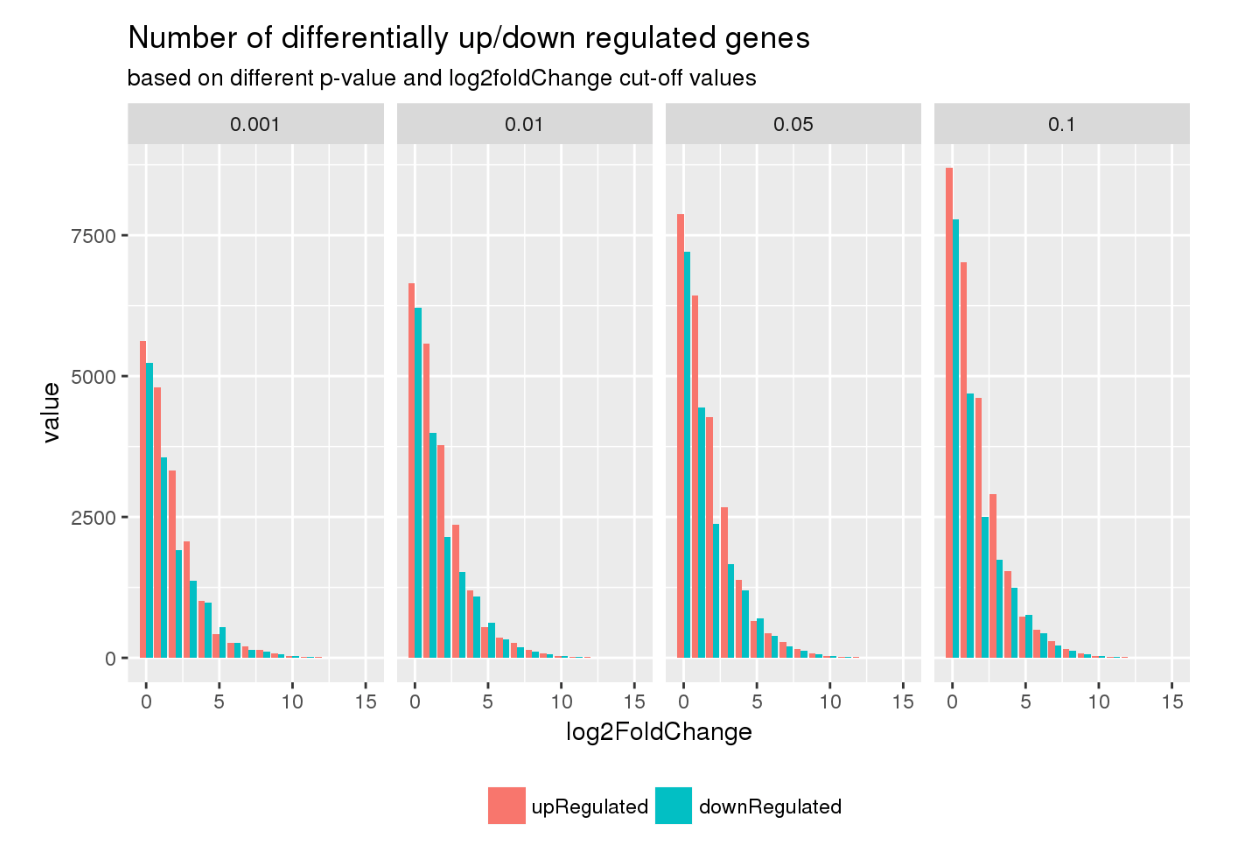
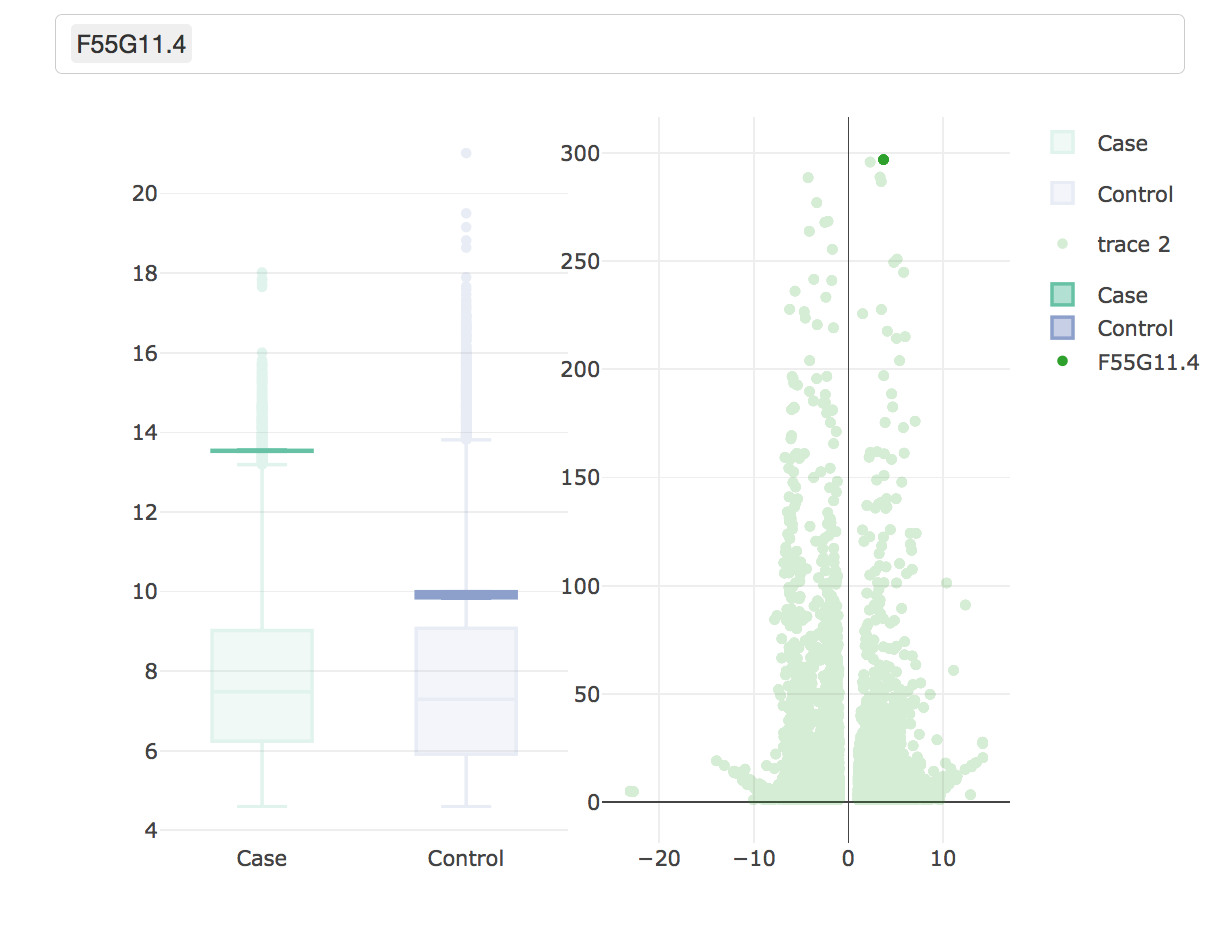
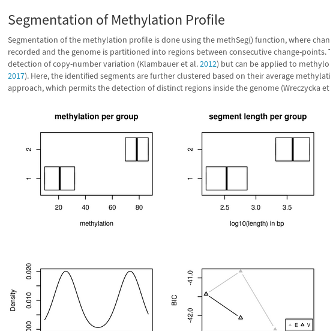

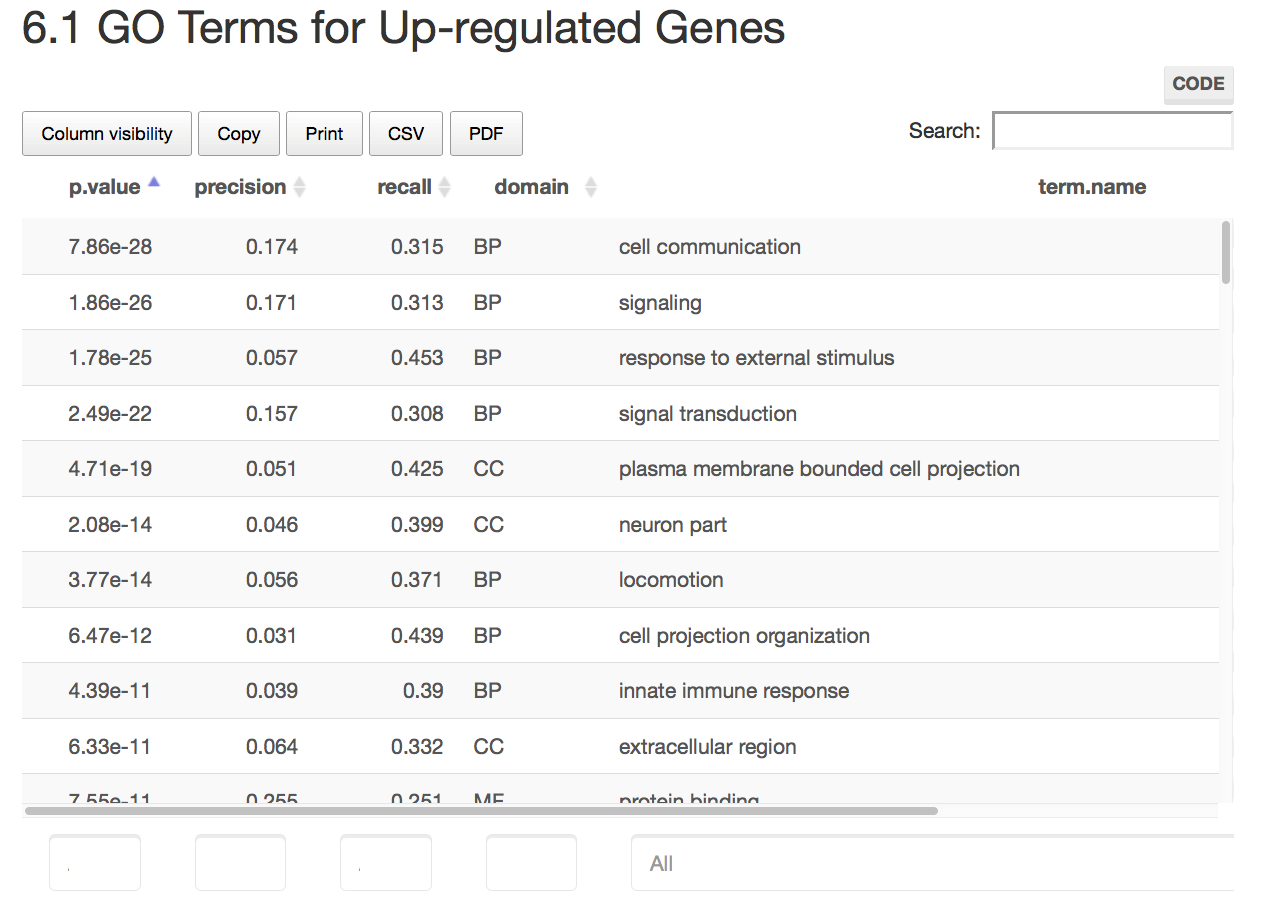
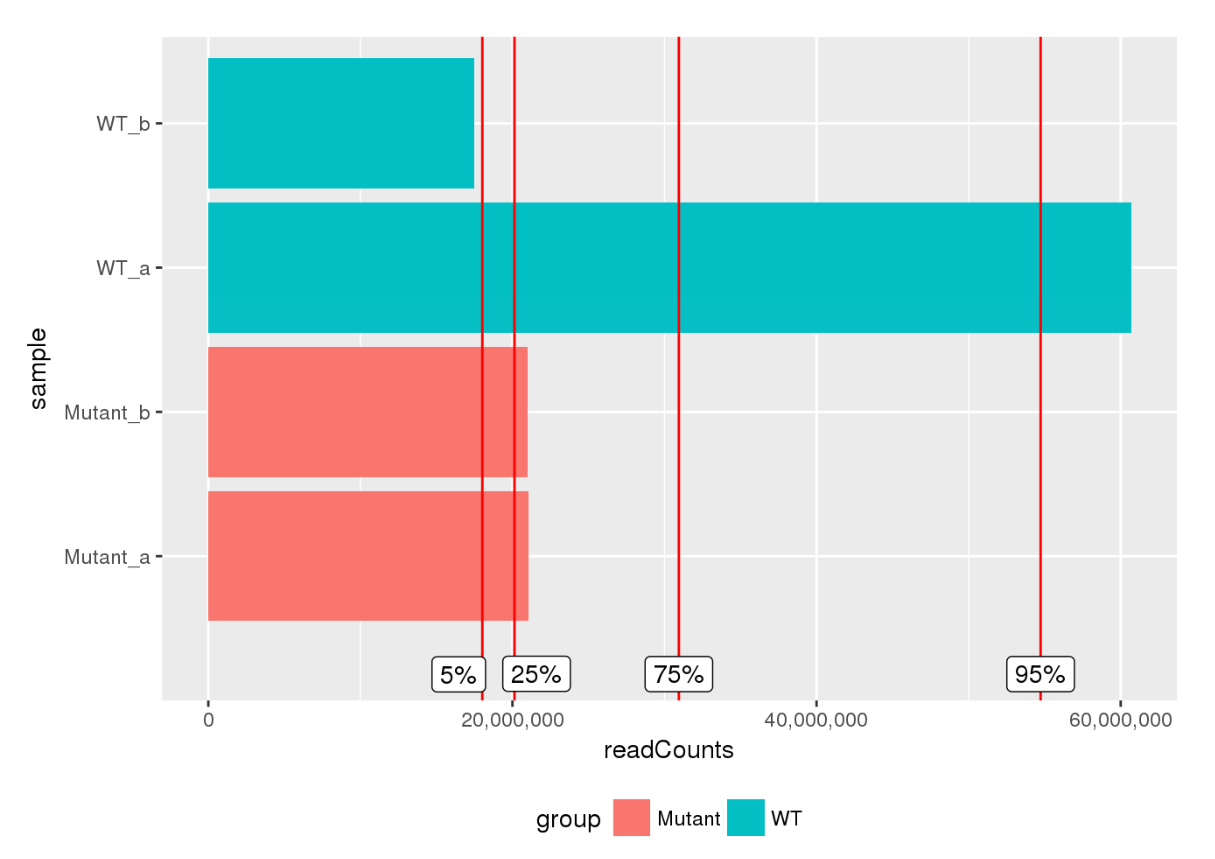
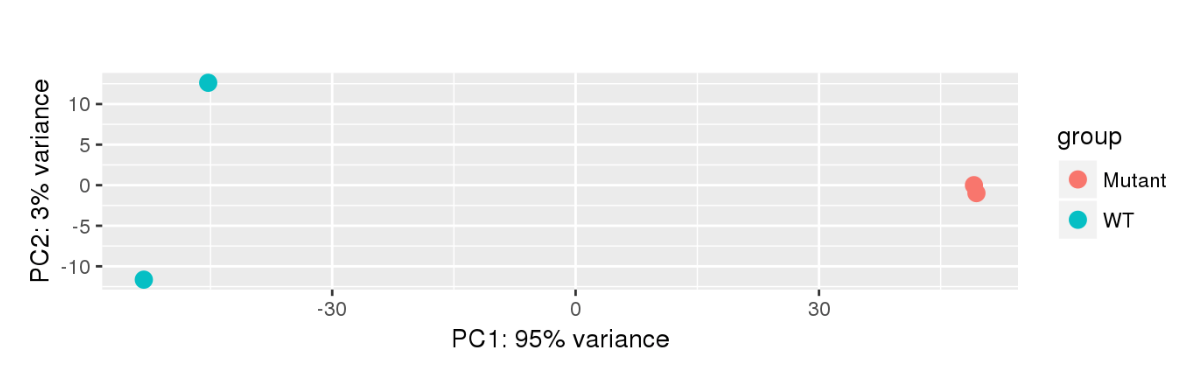
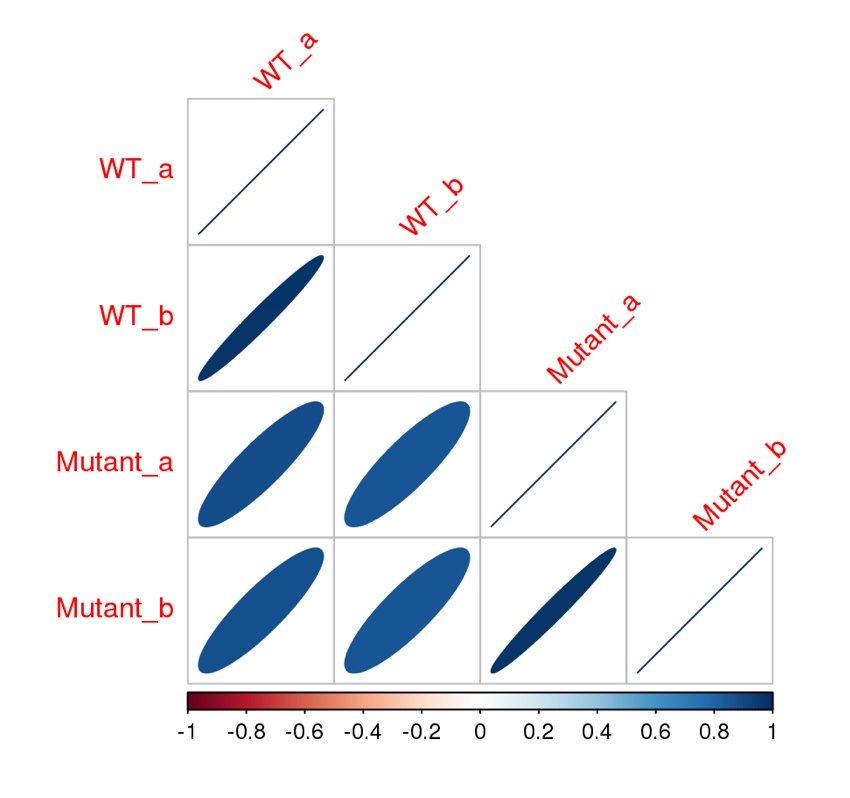
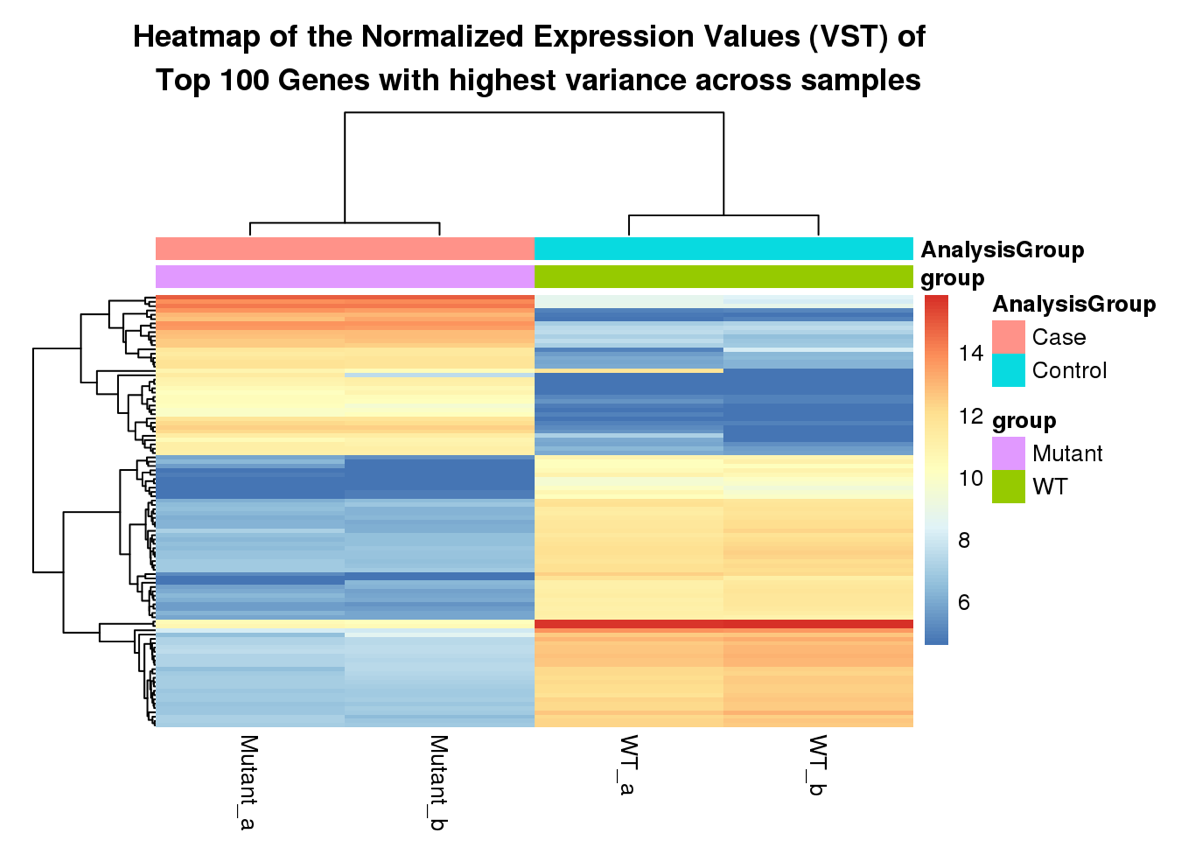
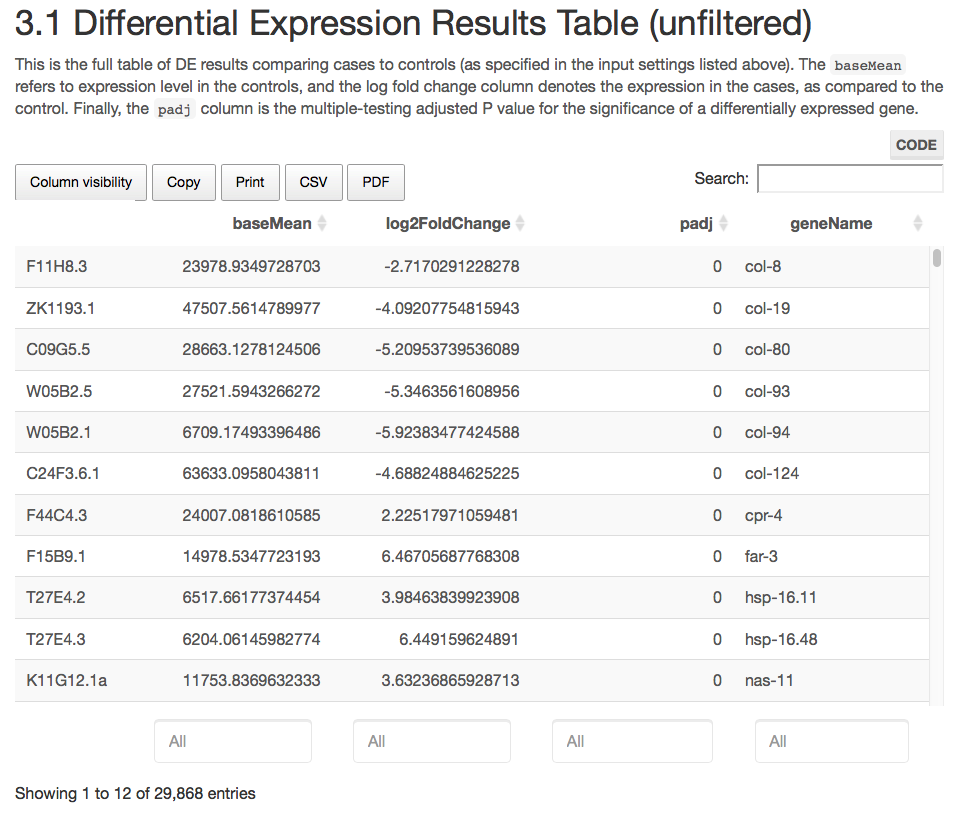
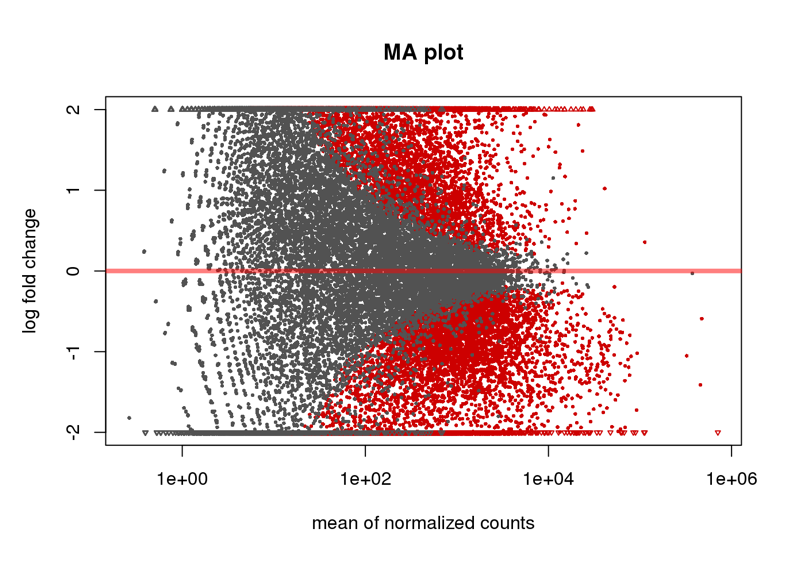
To run PiGx on your experimental data you only need to follow these three steps:
Describe your samples. Describe all your samples in a CSV file
sample_sheet.csv. To generate a sample sheet template for any
pipeline run this command:
pigx [pipeline] --init=sample-sheetTweak the default settings. Specify input and output
directories, and override defaults by providing a settings.yaml.
To generate a template for this file for any pipeline run this
command:
pigx [pipeline] --init=settingsRun the pipeline! Here's a simple example to run the RNAseq
pipeline when the sample sheet sample_sheet.csv is in the current
working directory:
pigx rnaseq -s settings.yamlThat's it! After some time you'll get a bunch of reports that you can view in your browser.
For more detailed information about each of the pipelines see the online documentation.
If you have further questions that are not answered in the documentation, or if you think you found an error in the pipelines, please send email to the user mailing list or use the web forum.
Pre-built binaries for PiGx are available through GNU Guix, the functional package manager for reproducible, user-controlled software management. Install the complete pipeline bundle with the following command:
guix install pigxPiGx is free software under the GNU General Public License (version 3 or, at your option, any later version). You can get the complete source code here. Your contributions are welcome!
Consider subscribing here to infrequent announcements regarding PiGx, such as release announcements.